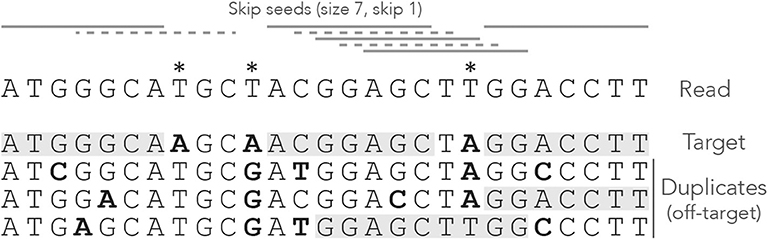
![PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a3f474fc4b86ec51d61e7fecab653f682693dd5f/2-TableI-1.png)
PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar
![PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a3f474fc4b86ec51d61e7fecab653f682693dd5f/3-Figure1-1.png)
PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar

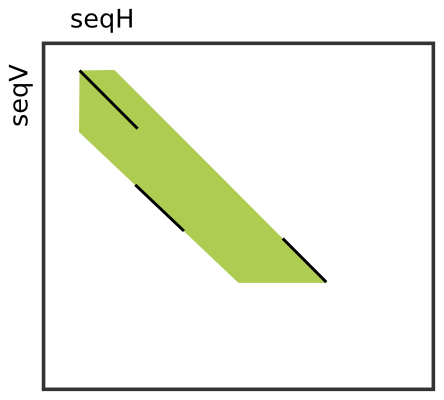
Comparison of traditional seed-and-extend alignment versus compressed... | Download Scientific Diagram
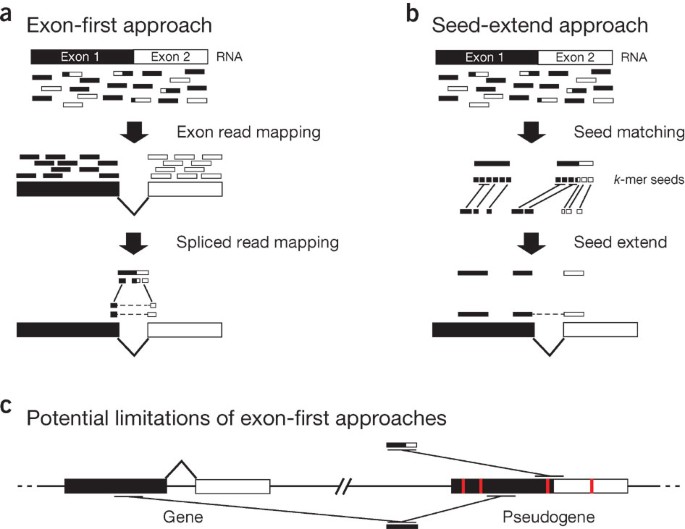
Computational methods for transcriptome annotation and quantification using RNA-seq | Nature Methods
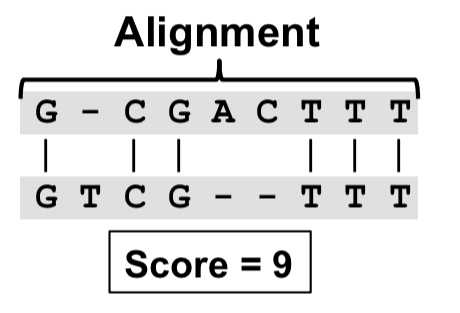
Darwin: a genomics co-processor provides up to 15,000x acceleration on long read assembly | the morning paper

Comparison of traditional seed-and-extend alignment versus compressed... | Download Scientific Diagram

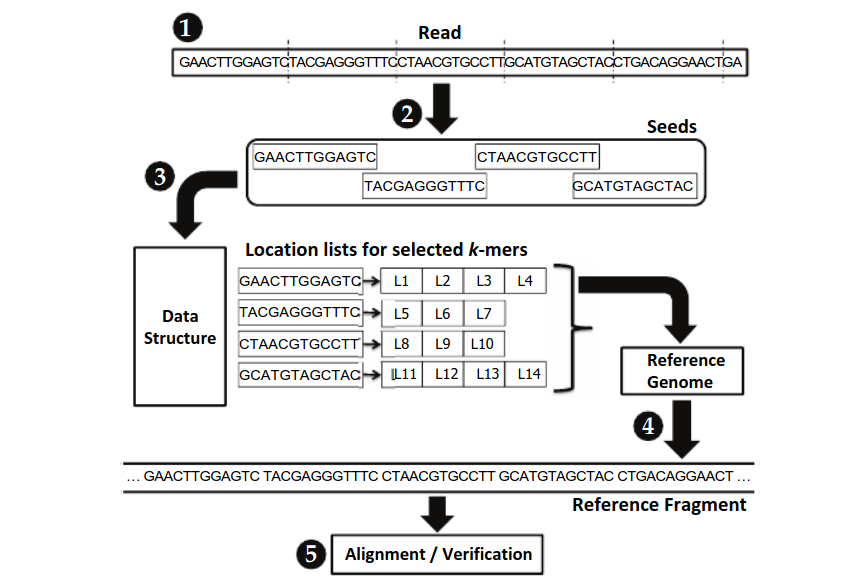
Seed-and-extend strategy. The alignment between the fragment of the... | Download Scientific Diagram

Seed-and-extend strategy. The alignment between the fragment of the... | Download Scientific Diagram
![PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a3f474fc4b86ec51d61e7fecab653f682693dd5f/3-Figure2-1.png)
PDF] A comparison of seed-and-extend techniques in modern DNA read alignment algorithms | Semantic Scholar
Power-Efficiency Analysis of Accelerated BWA-MEM Implementations on Heterogeneous Computing Platforms

Comparison of traditional seed-and-extend alignment versus compressed... | Download Scientific Diagram
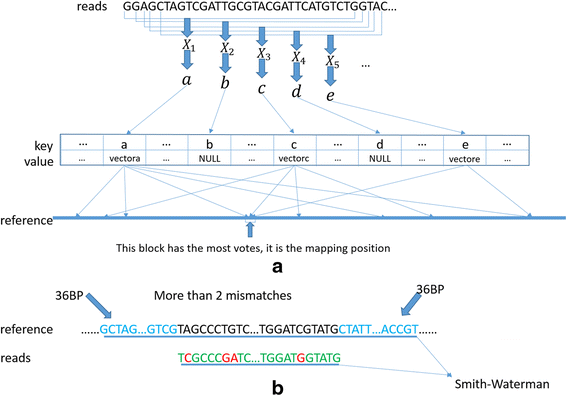
A fast read alignment method based on seed-and-vote for next generation sequencing | BMC Bioinformatics | Full Text
![PDF] OMBlast: alignment tool for optical mapping using a seed-and-extend approach | Semantic Scholar PDF] OMBlast: alignment tool for optical mapping using a seed-and-extend approach | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/7810149536dc5b06562a4d3d107adc7dfb075efa/3-Figure1-1.png)
PDF] OMBlast: alignment tool for optical mapping using a seed-and-extend approach | Semantic Scholar
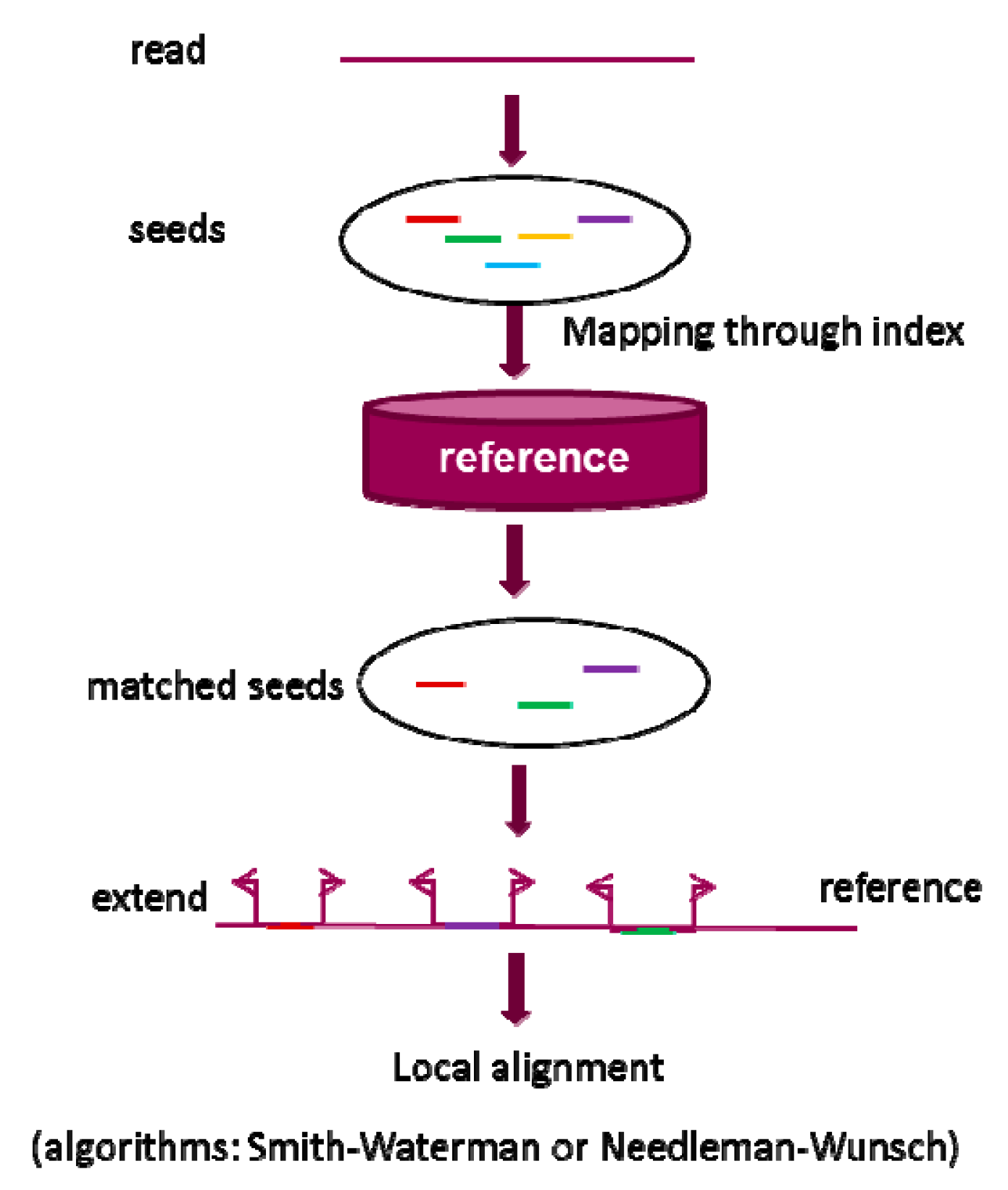
Pharmaceutics | Free Full-Text | Alignment of Short Reads: A Crucial Step for Application of Next-Generation Sequencing Data in Precision Medicine

Figure 1.2 from Accelerating the Understanding of Life's Code Through Better Algorithms and Hardware Design | Semantic Scholar
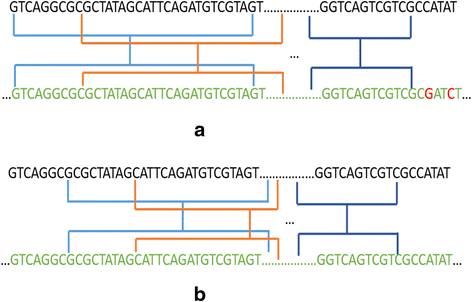
A fast read alignment method based on seed-and-vote for next generation sequencing | BMC Bioinformatics | Full Text