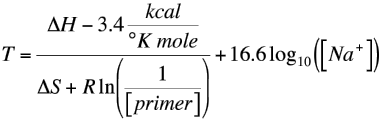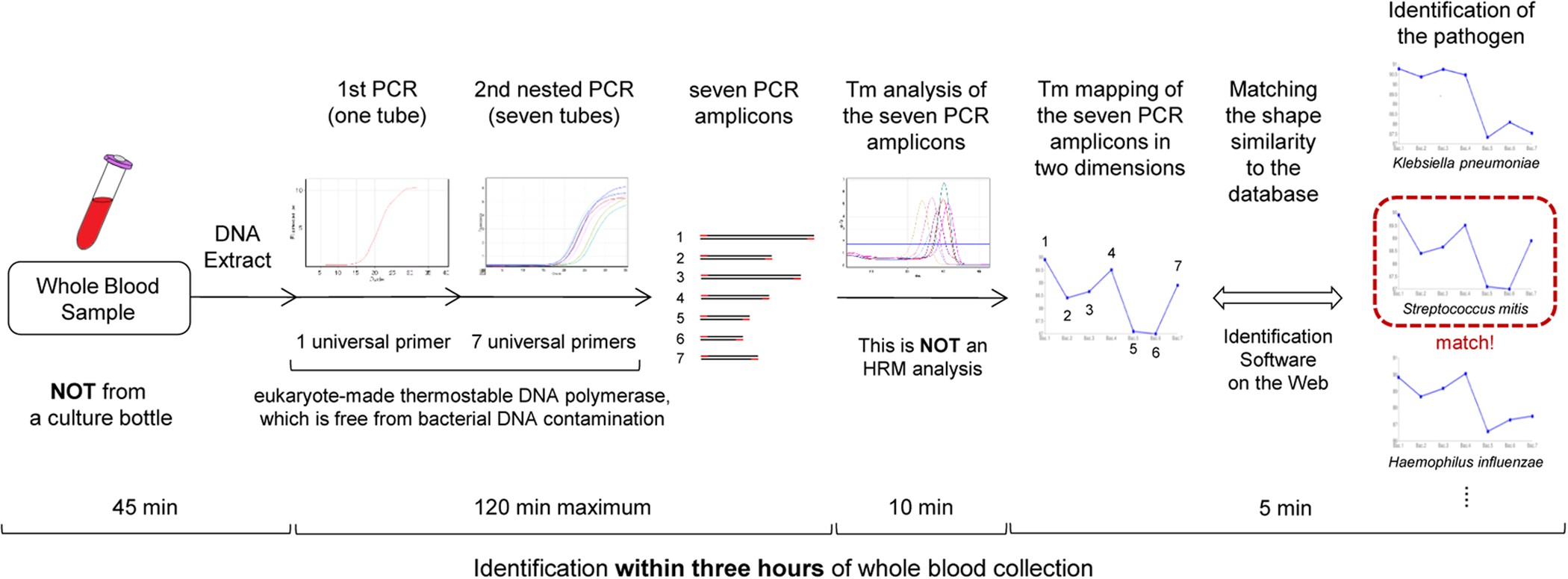
Melting Temperature Mapping Method: A Novel Method for Rapid Identification of Unknown Pathogenic Microorganisms within Three Hours of Sample Collection | Scientific Reports
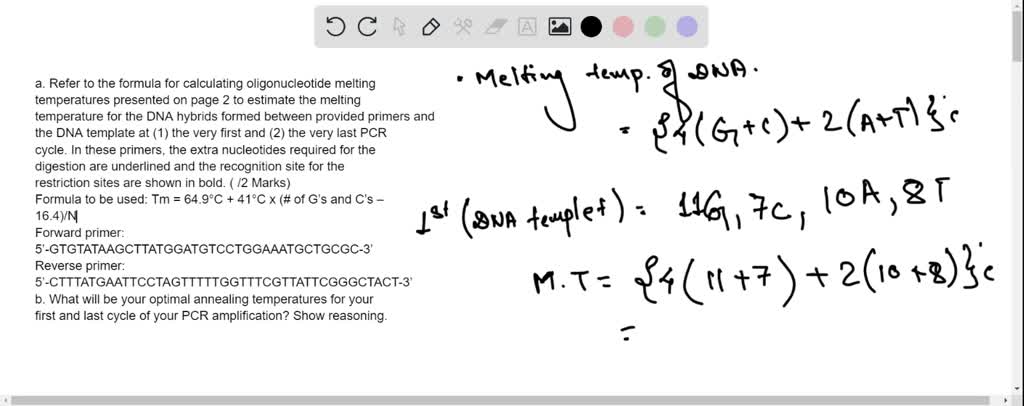
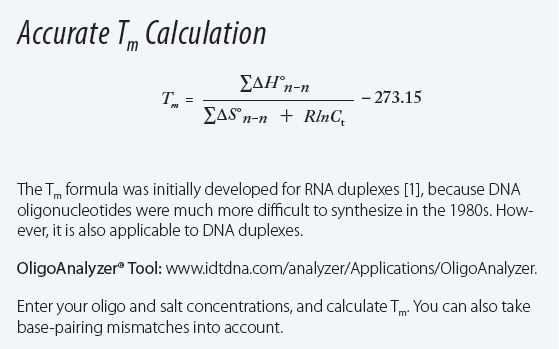
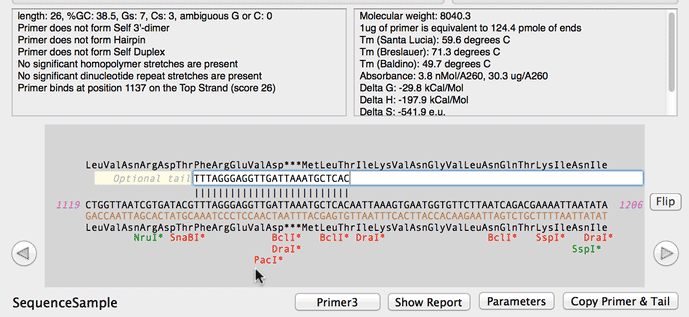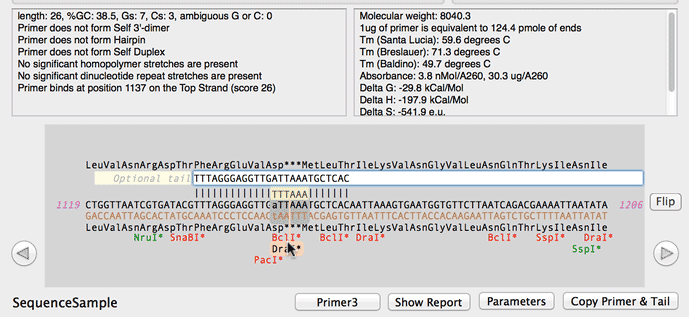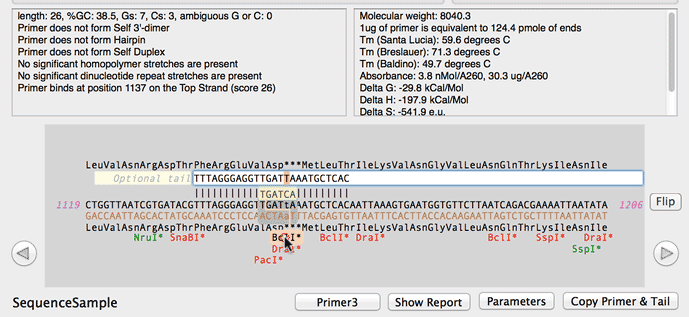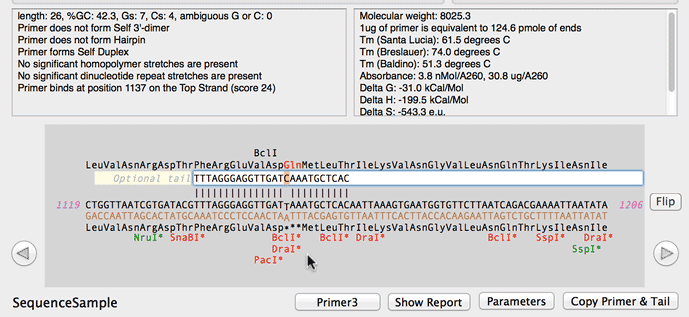
SOLVED: a. Refer to the formula for calculating oligonucleotide melting temperatures presented on page 2 to estimate the melting temperature for the DNA hybrids formed between provided primers and the DNA template

SOLVED: In this experiment; the same two original bookended primers will be used to amplify the ImrA sequence using PCR and the primary recombinant plasmid pAMG2S. The primers are: Forward: 5'-ATGGAAAGAGGTCCACAAATGGCCAAT-3' (Tm =

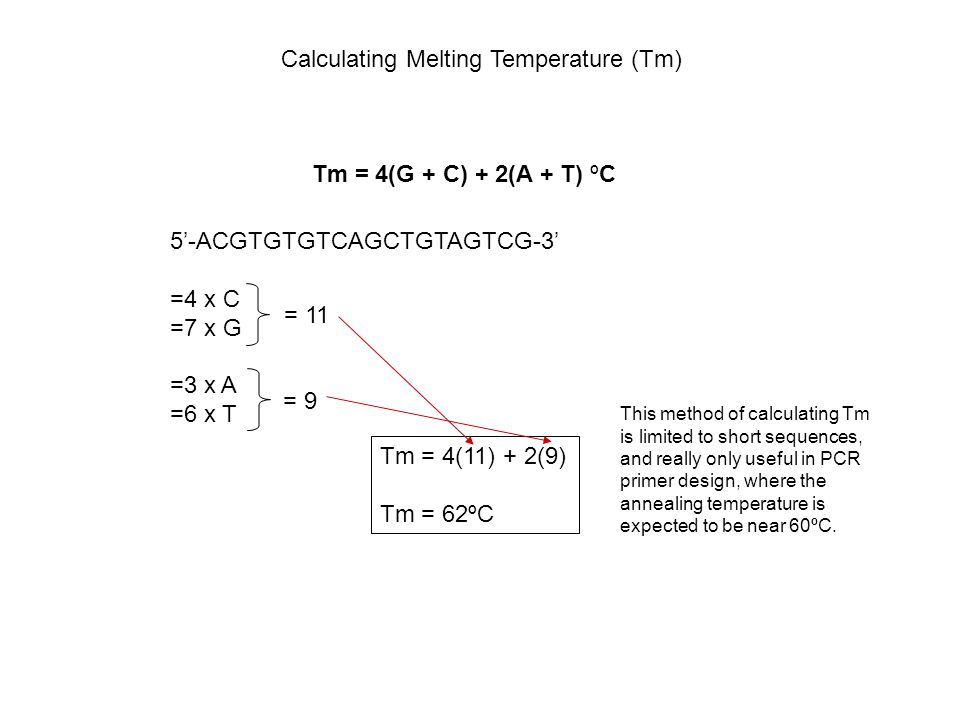
SOLVED: The annealing temperature of a DNA duplex is given by the formula: Tm 4(# of G-C basepairs) 2( # of A-T basepairs) Calculate the annealing temperature for the following primers: 5'ATGCCTATAGAATCGTC3'
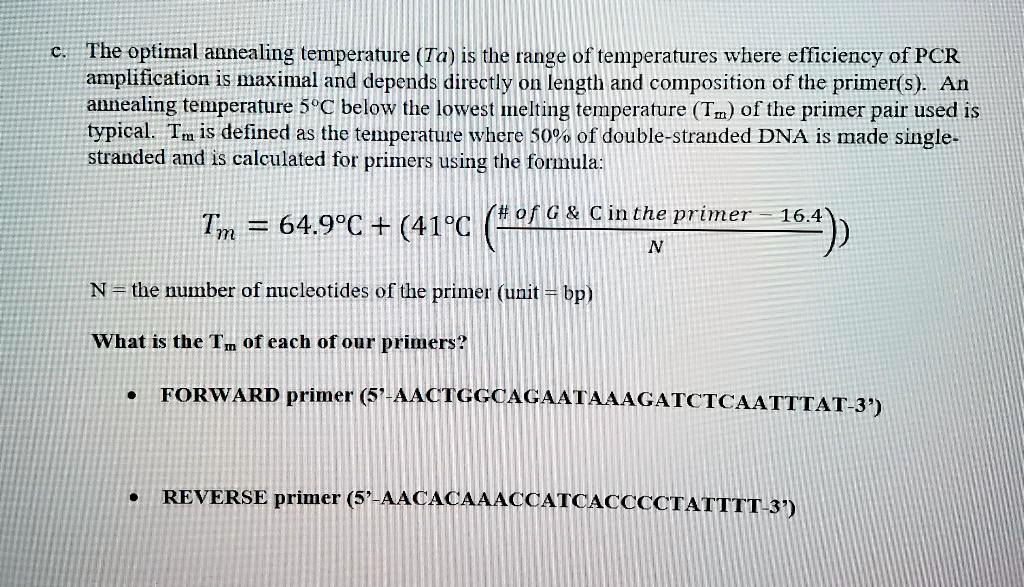

SOLVED: The optimal annealing temperature (Ta) is the range of temperatures where efficiency of PCR amplification is maximal and depends directly On iength and composition of the primer(s) An annealing temperature 5"€